| Identification |
|---|
| YMDB ID | YMDB00325 |
|---|
| Name | 2-trans,6-trans-farnesyl diphosphate |
|---|
| Species | Saccharomyces cerevisiae |
|---|
| Strain | Baker's yeast |
|---|
| Description | Farnesyl pyrophosphate, also known as farnesyl diphosphoric acid or farnesyl-PP, belongs to the class of organic compounds known as sesquiterpenoids. These are terpenes with three consecutive isoprene units. Farnesyl pyrophosphate is a very hydrophobic molecule, practically insoluble (in water), and relatively neutral. |
|---|
| Structure | |
|---|
| Synonyms | - (2E,6E)-farnesol diphosphate
- (2E,6E)-Farnesyl diphosphate
- (2E,6E)-Farnesyl pyrophosphate
- (all-E)-Farnesyl diphosphate
- (E,E)-Farnesyl diphosphate
- (E,E)-Farnesyl pyrophosphate
- 2-trans,6-trans-Farnesyl diphosphate
- 2-trans,6-trans-Farnesyl pyrophosphate
- all-trans-farnesyl diphosphate
- all-trans-Farnesyl pyrophosphate
- Farnesyl diphosphate
- Farnesyl pyrophosphate
- Farnesyl pyrophosphic acid
- Farnesyl-PP
- FPP
- fpp, farnesyl pyrophosphate
- omega,E,E-farnesyl diphosphate
- trans-Farnesyl pyrophosphate
- trans-trans-Farnesyl diphosphate
- trans-trans-Farnesyl pyrophosphate
- trans, trans-farnesyl diphosphate
- trans,trans-farnesyl diphosphate
- (2E,6E)-Farnesol diphosphoric acid
- (2E,6E)-Farnesyl diphosphoric acid
- (2E,6E)-Farnesyl pyrophosphoric acid
- (all-e)-Farnesyl diphosphoric acid
- (e,e)-Farnesyl pyrophosphoric acid
- 2-trans,6-trans-Farnesyl pyrophosphoric acid
- all-trans-Farnesyl pyrophosphoric acid
- Farnesyl diphosphoric acid
- trans,trans-Farnesyl diphosphoric acid
- trans-trans-Farnesyl diphosphoric acid
- 2-trans,6-trans-Farnesyl diphosphoric acid
- Farnesyl pyrophosphoric acid
- Farnesyl pyrophosphate, (e,e)-isomer
- Farnesyl pyrophosphate, (e,Z)-isomer
- Farnesyl pyrophosphate, (Z,e)-isomer
- Farnesyl pyrophosphate, (Z,Z)-isomer
- Farnesylpyrophosphate
|
|---|
| CAS number | 13058-04-3 |
|---|
| Weight | Average: 382.3261
Monoisotopic: 382.131026274 |
|---|
| InChI Key | VWFJDQUYCIWHTN-YFVJMOTDSA-N |
|---|
| InChI | InChI=1S/C15H28O7P2/c1-13(2)7-5-8-14(3)9-6-10-15(4)11-12-21-24(19,20)22-23(16,17)18/h7,9,11H,5-6,8,10,12H2,1-4H3,(H,19,20)(H2,16,17,18)/b14-9+,15-11+ |
|---|
| IUPAC Name | {[hydroxy({[(2E,6E)-3,7,11-trimethyldodeca-2,6,10-trien-1-yl]oxy})phosphoryl]oxy}phosphonic acid |
|---|
| Traditional IUPAC Name | farnesyl diphosphate |
|---|
| Chemical Formula | C15H28O7P2 |
|---|
| SMILES | CC(C)=CCC\C(C)=C\CC\C(C)=C\COP(O)(=O)OP(O)(O)=O |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as sesquiterpenoids. These are terpenes with three consecutive isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesquiterpenoids |
|---|
| Direct Parent | Sesquiterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Farsesane sesquiterpenoid
- Sesquiterpenoid
- Organic pyrophosphate
- Isoprenoid phosphate
- Monoalkyl phosphate
- Alkyl phosphate
- Phosphoric acid ester
- Organic phosphoric acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Charge | 0 |
|---|
| Melting point | Not Available |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Water Solubility | Not Available | PhysProp | | LogP | Not Available | PhysProp |
|
|---|
| Predicted Properties | |
|---|
| Biological Properties |
|---|
| Cellular Locations | |
|---|
| Organoleptic Properties | Not Available |
|---|
| SMPDB Pathways | | Cholesterol biosynthesis and metabolism CE(10:0) | PW002545 |   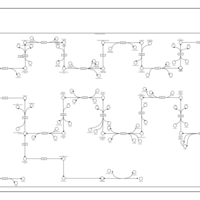 | | Cholesterol biosynthesis and metabolism CE(12:0) | PW002548 | 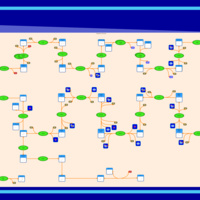 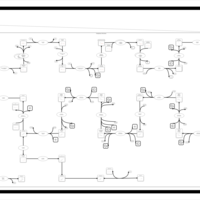 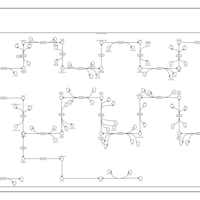 | | Cholesterol biosynthesis and metabolism CE(14:0) | PW002544 |    | | Cholesterol biosynthesis and metabolism CE(16:0) | PW002550 |  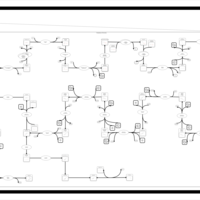 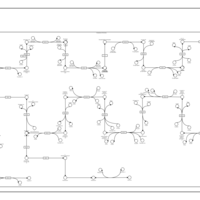 | | Cholesterol biosynthesis and metabolism CE(18:0) | PW002551 | 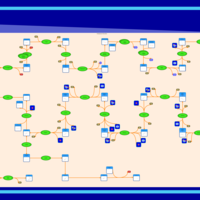 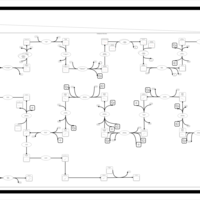  |
|
|---|
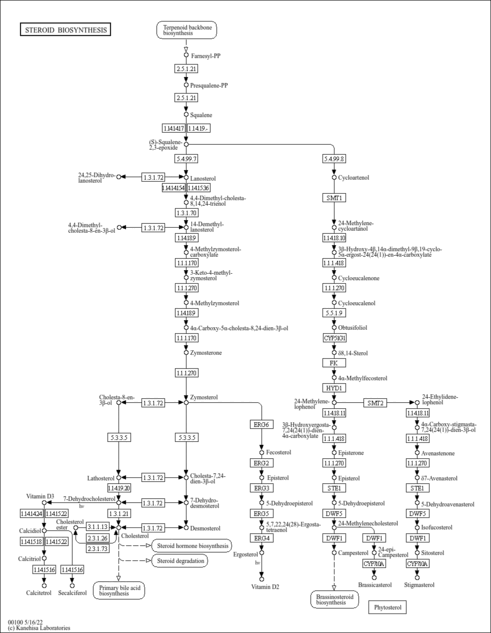
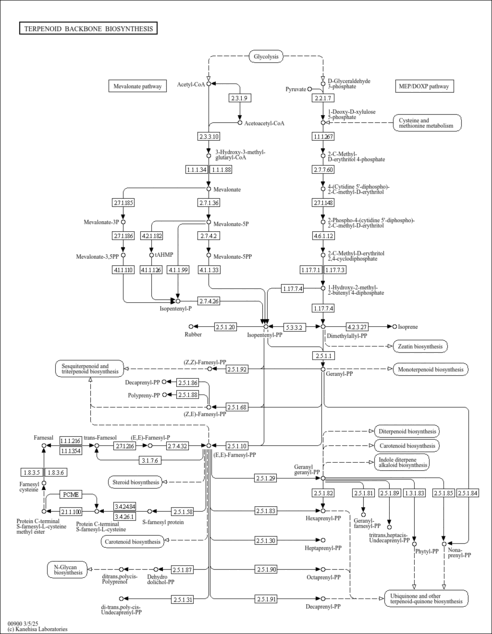
| KEGG Pathways | |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | |
|---|
| Concentrations |
|---|
| Intracellular Concentrations | Not Available |
|---|
| Extracellular Concentrations | Not Available |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-004j-7943000000-ab6d749700f510a94133 | JSpectraViewer | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | JSpectraViewer | | LC-MS/MS | LC-MS/MS Spectrum - n/a 26V, negative | splash10-014i-0090000000-09efa5f4e481376eae3d | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - Orbitrap 11V, negative | splash10-001i-2009000000-b2358e50ee86b2ebccb5 | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - Orbitrap 14V, negative | splash10-004i-9004000000-b255d40b3beef0ddd16f | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - Orbitrap 16V, negative | splash10-004i-9001000000-64cd48aa93040d775977 | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - Orbitrap 19V, negative | splash10-004i-9000000000-84a4e8d938e03661128e | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - Orbitrap 25V, negative | splash10-004i-9000000000-eb2e768f7c19af669f74 | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - n/a 26V, negative | splash10-03di-0209000000-651c59dc3e732ea880d7 | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - n/a 26V, negative | splash10-0a4i-0900000000-ae4be9bde4ed56cfa99b | JSpectraViewer | MoNA | | LC-MS/MS | LC-MS/MS Spectrum - n/a 26V, positive | splash10-0002-0294000000-7019f71a6e4f542dddae | JSpectraViewer | MoNA | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0pc0-1469000000-e3fd27c0418d0977f84e | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-6791000000-1ca128d2b96287a5b2f1 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0q29-9820000000-3277fbdf16e288ab1142 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0409000000-f8bbf786ee9d33cb48d4 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9501000000-8d060d3ceac94de45b8d | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-a265a369e6802359a7df | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0009000000-ea5bda9906940e9694b2 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-2409000000-4b044bb59695723b85bd | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-056r-9600000000-a090357e0efeb6192936 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0019000000-66a0a5510c1d34aea230 | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-114i-0394000000-89ab3170c9695823fb2f | JSpectraViewer | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-05rs-6900000000-7b5163a81d3cbdf1feb4 | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 13C NMR Spectrum | Not Available | JSpectraViewer | | 1D NMR | 1H NMR Spectrum | Not Available | JSpectraViewer |
|
|---|
| References |
|---|
| References: | - UniProt Consortium (2011). "Ongoing and future developments at the Universal Protein Resource." Nucleic Acids Res 39:D214-D219.21051339
- Scheer, M., Grote, A., Chang, A., Schomburg, I., Munaretto, C., Rother, M., Sohngen, C., Stelzer, M., Thiele, J., Schomburg, D. (2011). "BRENDA, the enzyme information system in 2011." Nucleic Acids Res 39:D670-D676.21062828
- Herrgard, M. J., Swainston, N., Dobson, P., Dunn, W. B., Arga, K. Y., Arvas, M., Bluthgen, N., Borger, S., Costenoble, R., Heinemann, M., Hucka, M., Le Novere, N., Li, P., Liebermeister, W., Mo, M. L., Oliveira, A. P., Petranovic, D., Pettifer, S., Simeonidis, E., Smallbone, K., Spasic, I., Weichart, D., Brent, R., Broomhead, D. S., Westerhoff, H. V., Kirdar, B., Penttila, M., Klipp, E., Palsson, B. O., Sauer, U., Oliver, S. G., Mendes, P., Nielsen, J., Kell, D. B. (2008). "A consensus yeast metabolic network reconstruction obtained from a community approach to systems biology." Nat Biotechnol 26:1155-1160.18846089
|
|---|
| Synthesis Reference: | Not Available |
|---|
| External Links: | |
|---|