| Identification |
|---|
| YMDB ID | YMDB12708 |
|---|
| Name | DG(12:0/12:0/0:0) |
|---|
| Species | Saccharomyces cerevisiae |
|---|
| Strain | Brewer's yeast |
|---|
| Description | DG(12:0/12:0/0:0) belongs to the family of Diacylglycerols. These are glycerolipids lipids containing a common glycerol backbone to which at least one fatty acyl group is esterified. DG(12:0/12:0/0:0) is also a substrate of diacylglycerol kinase. It is involved in the phospholipid metabolic pathway. |
|---|
| Structure | |
|---|
| Synonyms | - 2,3-Dilaurin
- 2,3-Dilauroyl-sn-glycerol
- DG 0:0/12:0/12:0
- 2,3-Didodecanoyl-sn-glycerol
|
|---|
| CAS number | Not Available |
|---|
| Weight | Average: 456.708
Monoisotopic: 456.381474774 |
|---|
| InChI Key | OQQOAWVKVDAJOI-RUZDIDTESA-N |
|---|
| InChI | InChI=1S/C27H52O5/c1-3-5-7-9-11-13-15-17-19-21-26(29)31-24-25(23-28)32-27(30)22-20-18-16-14-12-10-8-6-4-2/h25,28H,3-24H2,1-2H3/t25-/m1/s1 |
|---|
| IUPAC Name | (2R)-1-(dodecanoyloxy)-3-hydroxypropan-2-yl dodecanoate |
|---|
| Traditional IUPAC Name | (2R)-1-(dodecanoyloxy)-3-hydroxypropan-2-yl dodecanoate |
|---|
| Chemical Formula | C27H52O5 |
|---|
| SMILES | [H][C@@](CO)(COC(=O)CCCCCCCCCCC)OC(=O)CCCCCCCCCCC |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 1,2-diacylglycerols. These are diacylglycerols containing a glycerol acylated at positions 1 and 2. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Glycerolipids |
|---|
| Sub Class | Diradylglycerols |
|---|
| Direct Parent | 1,2-diacylglycerols |
|---|
| Alternative Parents | |
|---|
| Substituents | - 1,2-acyl-sn-glycerol
- Fatty acid ester
- Fatty acyl
- Dicarboxylic acid or derivatives
- Carboxylic acid ester
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Primary alcohol
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Charge | 0 |
|---|
| Melting point | Not Available |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Water Solubility | Not Available | PhysProp | | LogP | Not Available | PhysProp |
|
|---|
| Predicted Properties | |
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Endoplasmic reticulum
|
|---|
| Organoleptic Properties | Not Available |
|---|
| SMPDB Pathways | | Triacylglycerol metabolism TG(12:0/12:0/12:0) | PW007581 | 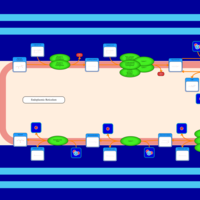 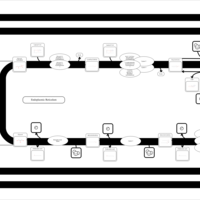 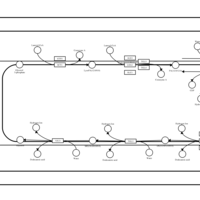 | | Triacylglycerol metabolism TG(12:0/12:0/22:1(13Z)) | PW007718 | 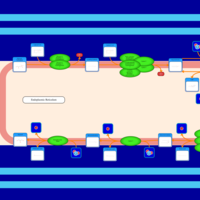 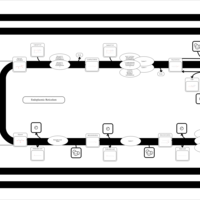 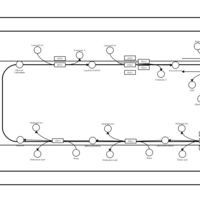 | | Triacylglycerol metabolism TG(12:0/12:0/25:0) | PW007763 | 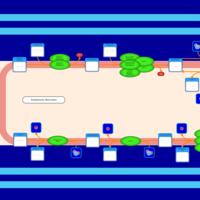 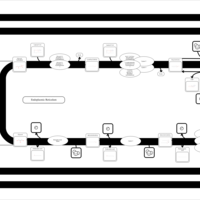 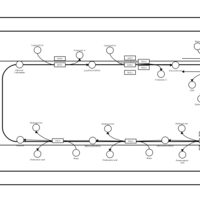 | | Triacylglycerol metabolism TG(12:0/12:0/27:0) | PW007787 |  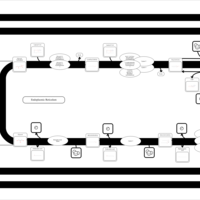 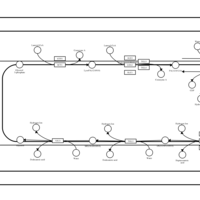 | | Triacylglycerol metabolism TG(12:0/12:0/29:0) | PW007815 | 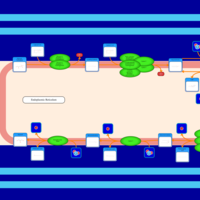 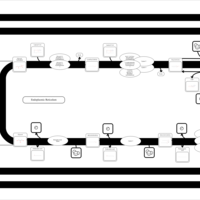 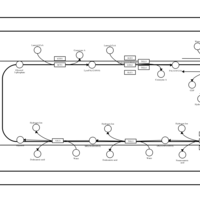 |
|
|---|
| KEGG Pathways | Not Available |
|---|
| SMPDB Reactions | Not Available |
|---|
| KEGG Reactions | Not Available |
|---|
| Concentrations |
|---|
| Intracellular Concentrations | Not Available |
|---|
| Extracellular Concentrations | Not Available |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-08fr-0090400000-7ee5f08309cde838a7ef | JSpectraViewer |
|
|---|
| References |
|---|
| References: | - Rattray JB, Schibeci A, Kidby DK. (1975). "Lipids of yeasts." Bacteriol Rev. 1975 Sep;39(3):197-231.240350
|
|---|
| Synthesis Reference: | Not Available |
|---|
| External Links: | | Resource | Link |
|---|
| CHEBI ID | 77390 | | HMDB ID | Not Available | | Pubchem Compound ID | 10456782 | | Kegg ID | Not Available | | ChemSpider ID | Not Available | | FOODB ID | Not Available | | Wikipedia ID | Not Available | | BioCyc ID | Not Available |
|
|---|
